Commutative Algebra Meets Data Science: A New Paradigm in Mathematical Artificial Intelligence
Commutative algebra is a longstanding pillar of pure mathematics that underpins subjects such as algebraic geometry and number theory and addresses fundamental structures like rings, ideals, and modules [2]. In the past, the field appeared to be quite removed from the practical concerns of data science. More recently, however, an unexpected transformation occurred — ideas from commutative algebra began to influence real-world data analysis; machine learning (ML); and even modeling efforts in biology, chemistry, and materials science.
The Emergence of Persistent Commutative Algebra
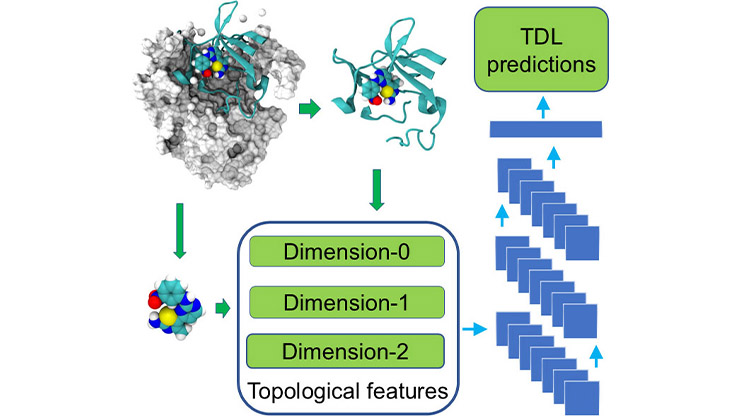
A burgeoning discipline called persistent commutative algebra (PCA) applies algebraic concepts in a multiscale, data-driven setting [8]. Much like topological data analysis (TDA) and persistent homology [1], PCA extracts structural “signatures” that remain stable across multiple scales or levels of resolution. But unlike TDA—which focuses primarily on topological invariants such as loops and voids—PCA captures algebraic relationships, constraints, and interactive patterns that are often invisible to topological, geometric, or statistical methods [4].
PCA is both mathematically elegant and practically powerful, with applications in genomics, molecular binding prediction, protein-nucleic acid interactions, and other complex biological systems. These developments suggest that the field may soon become a vital component of rational learning, which is guided by interpretable mathematics rather than opaque computation.
![<strong>Figure 1.</strong> Persistent commutative algebra analysis on a regular octagon. <strong>1a.</strong> Filtration of the octagon. <strong>1b.</strong> Persistent facet ideals and persistent graded Betti curves. 1c. Persistent
\(f\)-vectors and persistent \(h\)-vectors. Figure courtesy of Yiming Ren and adapted from [6].](/media/abefttbw/figure1.jpg)
Why Commutative Algebra, and Why Now?
While classical ML—particularly deep learning—has made extraordinary progress, it often lacks interpretability and structural awareness. Many existing models operate as black boxes, offering minimal insight into their behavior under perturbations or the rationale of their predictions. A key obstacle in the development of the next generation of artificial intelligence (AI)—specifically world models—is the lack of physical and structural insights in current large language models. As scientific datasets grow progressively more complex, these limitations become increasingly problematic.
Commutative algebra offers a different perspective by providing tools that capture algebraic relations, combinatorial dependencies, topological invariants, and geometric constraints. These mathematical structures arise naturally in point clouds, graphs, directed graphs, hypergraphs, networks, sequences, and other forms of data. PCA adapts the structures to data science by examining the emergence, evolution, and persistence of patterns across scales [8], yielding a multiscale viewpoint that parallels the success of TDA but extends far beyond the capabilities of topology. Recent work affirms that PCA can use concepts such as persistent facet ideals, persistent graded Betti numbers, persistent \(f\)-vectors, and persistent \(h\)-vectors—mathematical invariants that record changing relationships in the data during filtration—to deliver combined interpretations of the data at hand [6] (see Figure 1).
Algebraic Signatures for Real Data
Instead of merely describing data by shape or distribution, PCA focuses on algebraic relations that remain meaningful across various levels of resolution. As such, it extracts multiscale algebraic “fingerprints” that summarize the interactions between data variables and are effective across the following domains.
Genomics: One of the earliest applications of PCA in computational biology is a framework called commutative algebra \(k\)-mer learning (CAKL) [7]. This method uses algebraic structures derived from \(k\)-mer sets to capture deep combinatorial patterns in genomic sequences. When tested on 11 benchmark datasets, CAKL outperformed several state-of-the-art methods during tasks like viral variant detection, phylogenetic tree analysis, and viral classification. This result demonstrates that PCA-based sequence analysis can be robust, scalable, and interpretable, even in the presence of noisy or highly variable genomic data.
Biomolecular interactions: PCA can also predict biomolecular interactions via commutative algebra ML (CAML) and commutative algebra neural networks (CANNs). These approaches utilize persistent Stanley-Reisner theory to generate algebraic signatures for molecular structures. In recent studies, CAML and CANNs have achieved competitive or superior performance when predicting protein-ligand binding affinities [3] and mutation-induced human diseases [9].
A new direction further builds on this idea via graded Betti number learning (GBNL) [10], which uses primary sequence data to directly predict protein-nucleic acid binding affinities. By combining PCA with modern sequence embeddings, GBNL offers interpretability and strong predictive performance without requiring three-dimensional structural information.
Materials science: Category-specific commutative algebra (CSCA) is the first PCA implementation in materials science [5]. It utilizes chemically aware, multiscale algebraic invariants to model metal-organic frameworks. CSCA has achieved state-of-the-art predictive accuracy for gas adsorption properties, thus providing superior interpretability, stability, and data efficiency when compared to traditional geometric or graph-based methods. It introduces a rigorous new paradigm for the discovery and understanding of porous materials.
What Makes PCA Distinct?
PCA is unique in its ability to capture the evolution of algebraic relationships across scales, which reveals persistent combinatorial features. And unlike geometry-based methods, it exhibits natural compatibility with discrete, symbolic, and relational data and works seamlessly with sequences, graphs, and other non-Euclidean data types. Moreover, the unique algebraic invariants of PCA correspond to meaningful structural patterns and provide insights that are otherwise unavailable from black box neural networks.
PCA also enriches feature spaces with algebraic information, improving the performance and stability of downstream ML models and algebra-grounded neural networks. Finally, PCA-based models have already exceeded traditional methods in accuracy and robustness for a variety of application areas, from genomics and macromolecules to materials science. Collectively, these advantages establish the discipline as a promising mathematical foundation for data science.
The Road Ahead
Although the field of PCA is young, several factors are accelerating its growth. Persistent Stanley-Reisner theory serves as the theoretical foundation for data analysis; computational tools like Macaulay2 make algebraic computations tractable for medium- and large-scale datasets; and recent publications show strong empirical results and increasing interest across mathematics, computer science, and the physical sciences. Nonetheless, challenges remain. Broader adoption of the PCA framework will require user-friendly software packages, standardized benchmarks, and scalable algorithms for massive datasets; in particular, the computation of graded Betti numbers for large data remains an obstacle. Additionally, the interpretation of algebraic invariants in domain-specific contexts will be essential for scientists and engineers.
Despite these challenges, PCA brings a new language to data science — a language that is not centered on points or shapes, but on relationships, constraints, algebraic stability, and invariants. As datasets become more complex and interconnected over time, the demand for interpretable, mathematical AI methods will continue to grow. PCA is poised to complement existing tools—e.g., linear algebra, statistics, topology, and geometry—as a standard component of the modern data scientist’s toolkit. Its rising success suggests that it will play a key role in the next stage of rational learning: scientifically meaningful machine intelligence that is guided by profound mathematical structures.
References
[1] Carlsson, G. (2009). Topology and data. Bull. Amer. Math. Soc., 46(2), 255-308.
[2] Eisenbud, D. (2013). Commutative algebra with a view toward algebraic geometry. In Graduate texts in mathematics (Vol. 150). New York, NY: Springer.
[3] Feng, H., Suwayyid, F., Zia, M., Wee, J., Hozumi, Y., Chen, C.-L., & Wei, G.-W. (2025). CAML: Commutative algebra machine learning — a case study on protein-ligand binding affinity prediction. J. Chem. Inf. Model., 65(13), 6732-6743.
[4] Hu, C., Wang, Y., Xia, K., Ye, K., & Zhang, Y. (2025). Commutative algebra-enhanced topological data analysis. Preprint, arXiv:2504.09174.
[5] Khaemba, C.S., Feng, H., Chen, D., Chen, C.-L., & Wei, G.-W. (2026). Commutative algebra modeling in materials science — A case study on metal-organic frameworks (MOFs). J. Chem. Inf. Model.
[6] Ren, Y., & Wei, G.-W. (2025). Interpretability and representability of commutative algebra, algebraic topology, and topological spectral theory for real-world data. Adv. Intell. Discov., e202500207.
[7] Suwayyid, F., Hozumi, Y., Feng, H., Zia, M., Wee, J., & Wei, G.-W. (2025). CAKL: Commutative algebra \(k\)-mer learning of genomics. Preprint, arXiv:2508.09406.
[8] Suwayyid, F., & Wei, G.-W. (2026). Persistent Stanley-Reisner theory. Found. Data Sci., 8, 287-312.
[9] Wee, J., Suwayyid, F., Zia, M., Feng, H., Hozumi, Y., & Wei, G.-W. (2025). Commutative algebra neural network reveals genetic origins of diseases. Preprint, arXiv:2509.26566.
[10] Zia, M., Suwayyid, F., & Wei, G.-W. (2025). GBNL: Graded Betti number learning of complex biological data. Preprint, arXiv:2510.23187.
About the Author
Guo-Wei Wei
MSU Research Foundation Distinguished Professor, Michigan State University
Guo-Wei Wei is an MSU Research Foundation Distinguished Professor at Michigan State University. His research explores the mathematical foundations of bioscience and data science.
Related Reading
Stay Up-to-Date with Email Alerts
Sign up for our monthly newsletter and emails about other topics of your choosing.